WP1: Data Production
Tumors are heterogeneous with regions containing different genetic or epigenetic aberrations, which pose a great challenge for the prognostication of cancer. The biomarkers used to identify the aggressiveness of a tumor are not distributed evenly throughout the tumor, and by sampling only a minuscule amount of the tumor, there is a great risk of missing the cells that might lead to a patient’s death.
The experiments in this project have been designed such that multiple samples from each patient and tumor are examined with the goal of modelling the heterogeneity in prostate, colorectal, and lung cancer and analyzing the effect of sampling on the strength of the prognostic markers.
Work package 1 (WP1) has been responsible for the production of all samples to be analyzed in DoMore! The DoMore! project so far includes more than 66 000 samples of 2 886 tissue blocks from 7 028 patients.
Deliverables for WP1 were as follows: For each tissue block or biopsy included in the DoMore! project the lab was to produce 1-4 virtual slides, one DNA section and one monolayer.
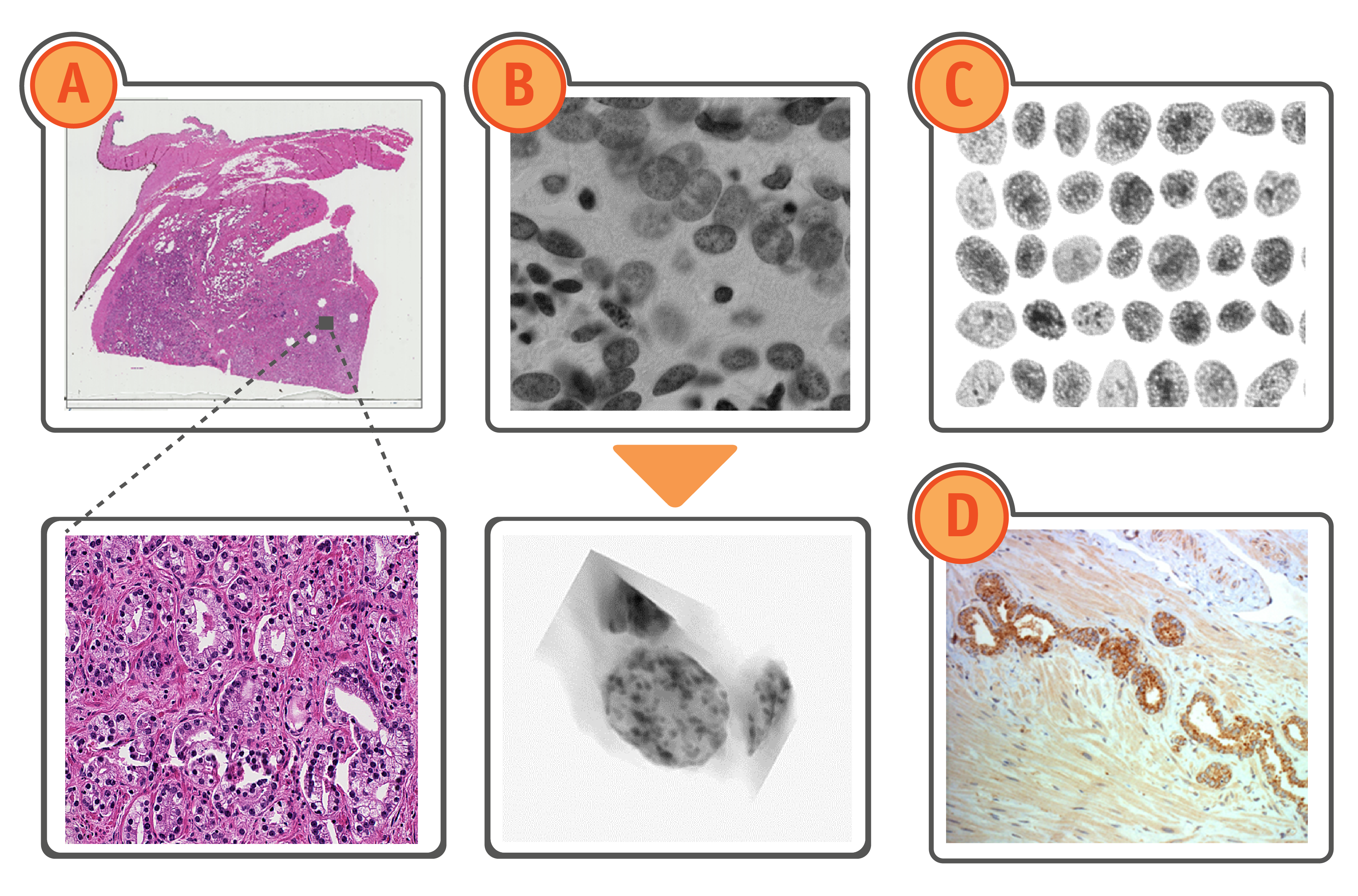 Image types: Virtual slide (A), DNA section (B), Monolayer (C) and gene expression shown with antibody binding to protein (D).
Image types: Virtual slide (A), DNA section (B), Monolayer (C) and gene expression shown with antibody binding to protein (D).
Virtual slides
A virtual slide is a scanned H&E stained 3µm tissue section, whereas the virtual slide production line in the laboratory also involves all tissue sectioning for all image types. Virtual slides are used in WP2 for Histotyping (tumor grading), WP3 for tumor delineation and in WP4 for quantification of stroma, Gleason grading, mitotic index and microtracking (co-analysis of different features on a cell-to-cell basis).
Each block is sectioned, obtaining (at least, but not limited to): one 3µm section, two 5µm thick sections, and one 50µm section. The 3µm section is stained with Hematoxylin and Eosin (H&E), and then our in-house pathologist marks the tumor area in each section. Using the H&E as guidance, the same area on the paraffin block is marked, with a sharp tool. When sectioning the 50µm scroll, we only collect the area marked as the tumor area. This is followed by a new 3µm section (stained with H&E), which the pathologist uses as a control section to verify that there is still tumour tissue left.
In order to develop methods that are invariant to imaging devices, all slides are scanned using scanners from at least two major vendors (Leica, Hamamatsu). Also, for the tumor delineation project in WP3, we needed tissue sections both with the pathologists marking of tumor area, as well as the same tissue section without any tumor marking. Based on this, we have so far produced 59 893 individual scanned H&E sections (Virtual slides) in DoMore!
DNA section
DNA sections are used in WP3 for automatic segmentation of cell nuclei, and in WP4for translation of DNA ploidy and Nucleotyping analysis from monolayers to tissue sections.
5µm sections are first stained with H&E and scanned, then de-stained and re-stained with Feulgen, a DNA specific stain, before it is scanned again. Our in-house pathologist marks the tumor area on the H&E section, the H&E and Feulgen stained sections are then overlaid using the marked tumor area on the H&E section to guide the measurement of DNA in the Feulgen stained section. The final measurements are initially performed on an automatic microscope system developed at the institute. We are currently also exploring the possibilities of replacing these microscope systems by scanners.
Monolayer
Monolayers, slides with isolated cell nuclei, are used for DNA ploidy and Nucleotyping analysis (see WP4).
From a 50µm thick tissue scroll cell nuclei are isolated and spun onto a glass slide. The nuclei are then stained with the DNA-specific Feulgen stain, and measured on an automatic microscope system developed at the institute.
Oslo, 15th February 2019
Navigate to the other Work Package descriptions: